About
Find Out More About AroCageDB
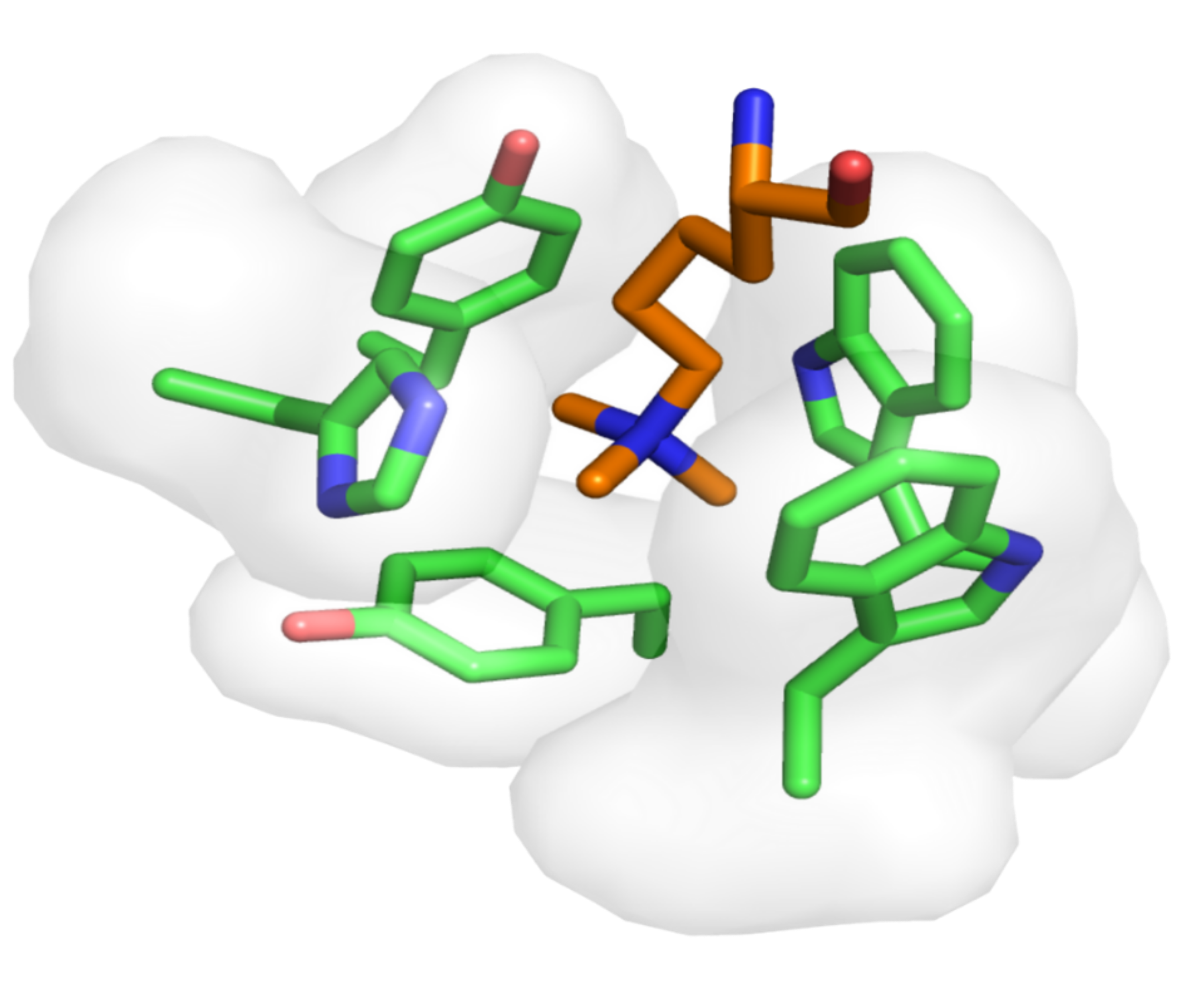
This database is a comprehensive, curated collection of aromatic cage structures from the Protein Data Bank ( https://www.rcsb.org/ ), analyzed for protein, structure, binding pocket, and ligand recognition studies. It was developed by the Pharmaceutical Bioinformatics group at the Institute for Pharmaceutical Sciences of the University of Freiburg (Germany). The database will be updated yearly in order to include more entries upon the deposition of new 3D models in the PDB.
For questions, please contact Prof. Dr. Stefan Günther (stefan.guenther@pharmazie.uni-freiburg.de).
Reference
Li J, Moumbock AFA, Qaseem A, Xu Q, Feng Y, Wang D, Günther S. AroCageDB: A Web-Based Resource for Aromatic Cage Binding Sites and Their Intrinsic Ligands. J Chem Inf Model. 2021 Nov 22;61(11):5327-5330. doi: 10.1021/acs.jcim.1c00927. Epub 2021 Nov 5. PMID: 34738791.Last Update: January 2021
1,636
Complexes
487
Proteins
818
Small molecule ligands
72
Peptide ligands
154
Organisms
Search
Explore our Database
Search with either PDB ID, protein name, uniprot ID, protein sequence or HET ID to inspect geometric properties of a specific aromatic cage, or to obtain lists of structures of a specific protein target, lists of homologous aromatic cage containing proteins, lists of proteins which all bind to a specific ligand or lists of ligands which all bind to a specific protein target. Or, perform similarity, substructure and scaffold searches with user-defined structures.
Protein Name/Uniprot ID
Search with one protein name/uniprot ID to obtain list of structures of one specific protein target
Similarity/Substructure
Perform similarity/substructure search with user-defined structure to obtain list of ligands with similar (sub)structure
Scaffolds
Perform scaffold search with user-defined substructure to obtain list of ligands containing one specific scaffold
F.A.Q
Frequently Asked Questions
-
What is AroCageDB?
AroCageDB is a comprehensive, manually curated collection of aromatic cage structures from the Protein Data Bank (PDB, https://www.rcsb.org/), analyzed for protein structure, binding pocket, and ligand recognition studies.
-
What is an aromatic cage?
An aromatic cage, also known as aromatic box or hydrophobic box, is a structural motif lined by two to five closely packed aromatic amino acid residues (Phe, Tyr, Trp, or His) in a highly hydrophobic binding site, often supplemented by a proximal anionic Asp or Glu residue(s). In proteins possessing aromatic cage domains, the specificity of molecular recognition is principally driven by multivalent cation−π interactions formed at the contact interface between a ligand’s cationic center and the aromatic cage residues and, to a lesser extent, by hydrophobic contacts.
-
Where does AroCageDB's content comes from?
All structures were curated from the PDB (https://www.rcsb.org/).
-
What is shape complexity and stereochemical complexity?
The complexity of a small molecule can be rationalized by evaluating its saturation (shape complexity, Csp3/[Csp2 + Csp3]) or its total content in chiral carbon atoms (stereochemical or structural complexity, Cstereogenic/Ctotal). P. A. Clemons et al. PANS USA 2010, 107, 18787.
-
How pocket descriptors are calculated?
The geometric descriptors of aromatic binding sites, namely pocket depth, volume, hydrophobicity, and druggability score, were calculated with DoGSiteScorer, a grid-based program that uses a Difference of Gaussians algorithm for pocket detection and a support vector machine model for druggability assessment. Volkamer, A. et al. J. Chem. Inf. Model. 2010, 50 (11), 2041–2052; Volkamer, A. et al. J. Chem. Inf. Model. 2012, 52 (2), 360–372.
-
Why pocket descriptors on some complex cards are displayed as 'None'?
Pocket descriptor evaluation failed in a few cases for shallow, solvent-exposed binding sites (e.g. PDB ID: 6J2P; UniProt ID: SPP1_YEAST). Thus pocket descriptors for these complexes are displayed as 'None'.
-
Why some downloaded PyMOL session files (PSE) do not show pocket in mesh?
Pocket shown as mesh in PSE files are calculated with DoGSiteScorer, a grid-based program that uses a Difference of Gaussians algorithm for pocket detection. Pocket detection failed in a few cases for shallow, solvent-exposed binding sites (e.g. PDB ID: 6J2P; UniProt ID: SPP1_YEAST). Thus pocket in those PSE files are not displayed as mesh.
-
How pocket similarities are calculated?
We calculated the pocket similarity using the APoc program. APoc (Alignment of Pockets), is an efficient program for large-scale structural comparison of protein pockets. We used the default parameters in APoc, which gives p-values and z-scores for each binding site comparison. APoc Ref: Gao, M. & Skolnick, J.; Bioinformatics 29, 597604 (2013).
-
What are the different third party plugins used?
We have embedded commonly used third-party plugins into the website to increase the functionality, namely: PDBe Molstar (https://github.com/PDBeurope/pdbe-molstar/) and Michelanglo (https://doi.org/10.1093/bioinformatics/btaa104) for the visualization of protein-ligand complexes; the command-line tool Molconverter (Marvin 20.18.0, 2020, ChemAxon, https://chemaxon.com/)for the depiction of 2D ligand structures, ChemDoodle (https://www.chemdoodle.com/) for structure editing, and RDKit (https://www.rdkit.org/)for similarity-based structure search, and BLAST+ (https://ftp.ncbi.nlm.nih.gov/blast/executables/blast+/LATEST/) for sequence similarity.
-
Whom should I contact with questions or suggestions?
Prof. Dr. Stefan Günther (stefan.guenther[at]pharmazie.uni-freiburg.de)
-
How do I cite the database?
Please cite:
Li, J. et al. AroCageDB: A Web-based Resource For Aromatic Cage Binding Sites and Their Intrinsic Ligands (manuscript in preparation).
Contact
Contact Us
We would love to hear from you! Have comments? Suggestions? Corrections?
Send us an email at stefan.guenther[at]pharmazie.uni-freiburg.de or get more information about our group home(hyperlink)
Our Address
Hermann-Herder-Strasse 9, D-79104 Freiburg i. Br.
Email Us
stefan.guenther[at]pharmazie.uni-freiburg.de
Call Us
+49 761 203 4871